Ideas
No Invasion Or Migration, But Interaction: What This New Genetic Study Suggests About Prehistoric India
Aravindan Neelakandan
Feb 05, 2019, 11:36 AM | Updated 11:35 AM IST
Save & read from anywhere!
Bookmark stories for easy access on any device or the Swarajya app.
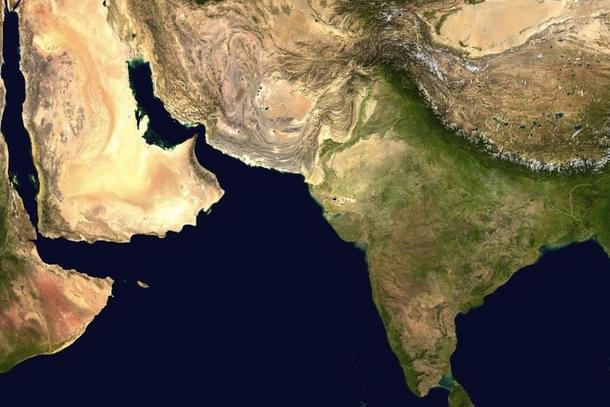

The year 2018 saw a series of articles in popular media that brought to public awareness the research that has been going on in molecular anthropology with relation to the peopling of ancient India.
You can also read this article in Hindi- न आक्रमण, न प्रवास- जानें भारतीयों के विषय में क्या कहता है अनुवांशिक अध्ययन
The new developments in ancient DNA (aDNA) recovery and examination have triggered this new-found interest. Often, these stories also come with a colonial baggage — the colonial narrative of Aryan and non-Aryan divide.
In India, the supposed Aryan migration, which happened almost 3,500 years before present (bp) has political ramifications. It is not uncommon to see those who write in public take advantage of the sensationalism surrounding any new discovery. Sometimes, even researchers may fall for it.
It is in such an environment that a new paper has come out which shows the very complex nature of ancient events involved in the peopling of India.
The study, 'The Genetic Ancestry of Modern Indus Valley Populations from Northwest India' (American Journal of Human Genetics) is done by Dr Ajai K Pathak of Estonian Biocentre, Institute of Genomics, University of Tartu and an international team.
The paper involves Toomas Kivisild, Mait Metspalu, Richard Villems — legendary names in the field of molecular anthropology and Dr Gyaneshwer Chaubey of Banaras Hindu University.
Dr Chaubey is an authority in the ancient genomic dynamics of the larger Indian land-mass called South Asia. The team consists of 17 researchers from world over.
The paper studies the “new genome wide genotype data for 45 modern individuals from four Northwest Indian populations” in the context of “several historic and prehistoric population movements between South Asia and West Eurasia”.
Of the groups studied, one community, the Ror, “stand out by being genetically more akin to populations living west of India”, the study says.

How is this important and how is this different from the other studies, which have now received a lot of media attention?
Dr Pathak answers the question by pointing out that “while most of the genetic studies from Northwest Indian part have been focusing on uniparental data that is used to trace paternal ancestry (Y chromosome) or to trace maternal ancestry (mtDNA), this study is based on the biparental genome data (ancestry contribution from both parents)”.
And that he says “gives a much clear picture together with filling the much-needed genetic hiatus from this important region”.
The Ror individuals that the study sampled were from within a 100 kilometre radius of Kurukshetra, whereas other sampled ethnic individuals too were from within 100 km to 400 km of the city. Sydney-based IITian Anurag Kadian, who was keen to understand the ancestral root of Ror population from Haryana, helped in the sample collection.
According to Dr Pathak, they have shown “a sort of genetic continuity between populations living west of the Indus valley and east of the Indus valley suggesting a higher mobility during prehistorical time in the Indus Valley region”.
The mobility might be because of a prolonged drought that another study had brought out earlier — a 900 years drought, which possibly triggered the collapse of the Indus Valley and migration of the Indus people toward the Gangetic Plain in search of favourable climate and water resources.
Another highlight of the study is the persistence of the genetic heterogeneity that one sees even today in the populations of the modern Indus region, which the scientists say might have come from ancient Indus time.
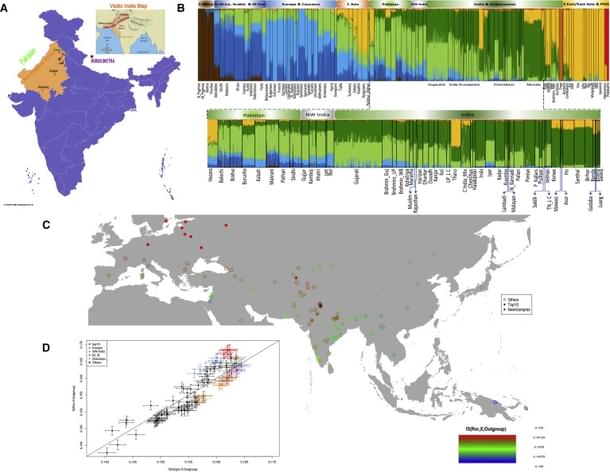
This is further corroborated by the earlier archaeological evidences which suggest that “Harappan culture must have integrated diverse regional and tribal features”, says Dr Pathak.
The most important aspect of the study is that by comparing ancient DNA worldwide, it shows the Ror and Jat communities as closer “to extant and ancient populations that live west of the India”.
But the mystery deepens further with the Ror and Jats having close affinity with “Central Asians and Bronze Age Steppe groups”, while also having “higher affinity with Anatolian Neolithic than Iranian Neolithic”.
This highlights them as a key population, which possibly “is very close to a root population that spread the Ancestral North Indian (ANI) component in South Asia due to high inward and outward mobility of the Indus people probably for trade”, explains Dr Pathak.
This interactionist model that Dr Pathak et al are uncovering, in a very elegant and ingenious study, is different from the templates of invasionist/migrationist models, which have been debated so far in this context.
Right now things are in a churn. At the same time, the admixtures from the steppe itself are categorised into two: one from the earlier to middle Bronze Age (Steppe_EMBA) and second from late to middle Bronze Age (steppe_MLBA).
One of the modellings used in the study also shows that “the NI_IE (North Indian Indo-European) group from the Gangetic Plain and Dravidian South Asians show a significant Steppe_EMBA component but not a Steppe_MLBA component” while the pre-historic and early historic South Asians from Swat Valley, Pakistan, “have a higher proportion of Steppe_MLBA than Steppe_EMBA”. So the puzzles are still far from being put together.
However, the interactionist model with bidirectional genetic admixtures is becoming more and more relevant.
Dr Chaubey, who is also part of the study, pointed out that “the intrusion into India at any period (after out of Africa) is minimum” and that “the genetic data doesn't fit with linguistic data”.

Further, while genetic marker R1a has been associated with steppe spread, Dr Chaubey has demonstrated in his recent presentation the in-situ origin of most frequent R1a branch - L657 (or M780).
However, the study has shown one thing very clearly — India and its cultural centres have always been home to diverse populations, which mixed together to form the people we know today as Indians.
Note: The original paper this article refers to can be found here:
Pathak et al. , 'The Genetic Ancestry of Modern Indus Valley Populations from Northwest India'.
Aravindan is a contributing editor at Swarajya.





